 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
|
Refuting Anon |
|
|
|
Note: This page has been recently updated and translated to Spanish by another webmaster. |
|
|
|
Y Chromosome and mtDNA haplogroups
An introduction to the use of Genetics in Anthropology |
|
|
|
Genetics brought a whole new perspective to the study of the human species. After analysing human DNA and comparing different sequences between individuals from different populations, scientists can fill in some of the blanks of the puzzle that Human History and Pre-History really are. By associating archaeological evidence and known historical events (such as the migrations of populations) with the variations in the presence of particular genetic sequences inside populations, the researchers are often able to trace the ancestry of a particular group of people and find previously unknown genetic relations between populations. |
|
|
|
 |
|
|
|
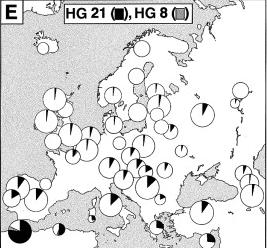
In the picture on the left, the populational frequencies of the Y chromosome sequence called HG21 are shown in black. As it can be seen, its frequency is at its highest in North Africa, and at southern Europe. One can imagine a North African origin to this sequence, eventually followed by a migration that dilutes its north African genes the more it migrates to the north. |
|
|
|
|
|
Two sources of DNA are of particular use to outline the history of Human populational migrations: the Y chromosome, and the mtDNA. |
|
|
|
The Y chromosome |
|
|
|
There are two types of variations (not connected to disease) within the DNA of a Y chromosome. The first one is a type of variation that changes quite rapidly, that partly allows us to distinguish different men in a population, and it sort of works like a personal chromosomal signature. The other kind of variation changes very slowly, so that there may well be large numbers of men with a similar type of Y chromosome. These DNA sequences that allow us to group the chromosomes in different "families" are called an haplogroup (HG). The haplogroup to which a Y chromosome belongs to provides in many cases a very clear evidence about its origin. |
|
|
|
 |
|
|
HG1 |
|
|
|
|
|
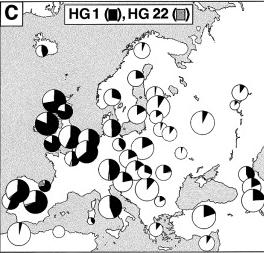
This is the most common haplogroup in Europe. The members of the HG1 chromosome family are regarded as the descendants of the Palaeolithic hunter-gatherers who arrived in Europe about 40.000 years ago. It is particularly common in Western Europe. HG1 is particularly high in Ireland (98,5%). The mutation defining HG1 has been dated at 23,000 YBP. |
|
|
|
|
|
|
HG2 |
|
|
|
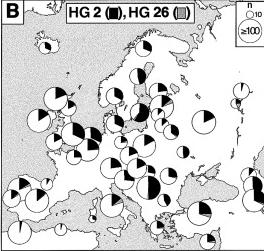
The members of HG2 are thought to be the descendants of the Neolithic waves of migrants that arrived about 10.000 years ago from the Southeast and that are credited with introducing agriculture into Europe. As they moved Northwest with their new technologies, they displaced the local hunter-gatherer populations (and thus diluted HG1).
Rousser et al. (2000) shares a different opinion. This study states that "chromosomes within this haplogroup are essentially undefined and are likely to consist of a set of different of discrete sublineages that themselves probably have greater geographic coherence. Consistent with this, HG2 (...) constitute the only high frequency lineage that does not show clinal variation. Because of this unimformativeness, HG2 will not be further considered here."
HG2 is most common in Southern and Central Europe. |
|
|
|
 |
|
|
|
|
|
HG3 |
|
|
|
 |
|
|
|
|
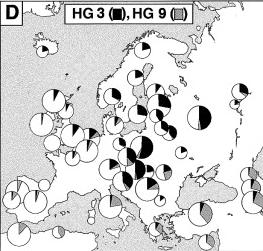
The forefather of HG3 may have been born in the Ukraine during the last Ice Age about 15.000 years ago. HG3 is seen more frequently on the Eastern side of Europe where it comprises approximately half the Russian, Polish and Slovakian Y chromosomes. There are several explanations for its frequencies in Europe. Cavalli-Sforza explained it with the Kurgan expansion from north of the Caspian Sea,dated to 7,000 YBP, while Rousser et al (2000) explained it through the spread of pastoralism, or with the East to West movements of such populations as the Scythians, Mongols, and Huns. |
|
|
|
|
|
|
HG8 |
|
|
|
HG8 is sub-Saharan in origin. It was only detected in Sardinian and French samples. |
|
|
|
HG9 |
|
|
|
HG9 reaches its highest frequencies (33%) in the Caucasus and in Anatolia. A simplistic interpretation is that HG9 chromosomes were carried in a major demographic expansion of agricultural migrants from the Near East. HG9 has been dated at 14,800 � 9,700 YBP. |
|
|
|
HG12 |
|
|
|
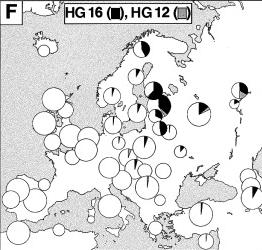
This chromosome is ancestral to HG16. It is not found in Western Europe. |
|
|
|
 |
|
|
HG16 |
|
|
|
|
|
HG16 is found at high frequency in the North, East of the Baltic Sea. With the exception of the Hungarians, all Uralic-speaking populations tested (Finnish, Estonians, Saami, and Mari) show a high frequency of HG16. |
|
|
|
|
|
|
 |
|
|
HG21 |
|
|
|
|
|
HG21 is North African. Within Europe, it reaches frequencies of some 27% of the Greek and Cypriot chromosomes, and in countries like Spain,Portugal, Sardinia, Italy, Turkey, and former Yugoslavia, it is between 10-20%. |
|
|
|
|
|
|
HG22 |
|
|
|
HG22 reaches appreciable numbers only in the French and the Basques. It was probably originally Iberian. |
|
|
|
MtDNA |
|
|
|
Introduction |
|
|
|
Mitochondrial DNA is a type of DNA that is passed from mother to child. During the event of fertilisation, the egg is fertilised by the male gamete. The human spermatozoid contributes with the Y chromosome and with 22 autosomal chromosomes. Everything else necessary for the development of the embryo comes from the egg. This includes, of course, the mitochondrial DNA. A mitochondria is a cellular organite responsible for the production of the energy that allows eukaryotic cells to work properly. It is of great use to the study of human populational migrations, since it is passed unchanged from mother to son and from mother to daughter. |
|
|
|
Scientists believe that there is about one change in the mtDNA (through mutation) every 10.000 years from the first modern human, referred to as mitochondrial Eve, who lived approximately 150.000 years ago in Africa. This allowed them to group mtDNA into families called haplogroups. An mtDNA haplogroup is therefore determined by polymorphisms [polymorphisms are the different possible genetic sequences for a given loci] that occurred thousands of years ago. Haplotypes are sub-clusters of haplogroups, and the polymorphisms that determine them are less prevalent, and a lot more recent. Most of the polymorphisms that determine haplogroups are continent specific. An haplogroup is coded with a capital letter (e.g. U) while its sub-clusters are coded with a number (e.g. U6). |
|
|
|
The earliest work with mtDNA started in the early 80s. Scientists either digested a small number of DNA samples using a large number of restriction enzymes [restriction enzymes are proteins that are able to cut DNA sequences in precise known locations], or they digested a large number of samples with a single enzyme. In either situation, if the DNA molecules are any different from each other, the enzymes will cut the DNA chains in different places (given the specificity of the enzymes), thus producing chopped sequences of different sizes. |
|
|
|
The analysis of mtDNA became an important tool for the study of human populational structure and history (Stoneking et al, 1993). Most phylogenetic analyses of mtDNA have been based on sequence variation in the first hypervariable segment of the control region (HVS-1) which is the most variable part of mtDNA. RFLP studies [an RFLP is a Restriction Fragment Length Polymorphism, and they are the chopped sequences obtained through the digestion of a DNA chain by restriction enzymes] of the mtDNA coding region have been used to classify mtDNA haplogroups. |
|
|
|
Mitochondrial DNA haplogroups |
|
|
|
As it was previously mentioned, the first modern human lived approximately 150.000 years ago in Africa. Soon afterwards, an early expansion of modern humans populated Africa and left its genetic fingerprints in mtDNA (the L1 haplogroup) found particularly in the KhoiSan (Bushmen) people today. This early expansion appears not to have left Africa, probably due to the harsh ice age climate and to the presence of the Man of Neanderthal in Eurasia. Tens of thousands of years later, an east African expansion (60-80.000 years ago) repopulated Africa (L2 and L3 types) and ultimately led to a migration out of Africa of at least one mtDNA subtype (L3a). |
|
|
|
L1 represents 29% of all African mtDNA's, while L2 encompasses some 34%. The remaining 37% belong to haplogroup L3, an heterogeneous group of 4 lineages. One of these lineages, defined by the loss of the Ddel site at the base pair 10394, represents only a few percent of the African mtDNAs, but appears to be the progenitor of roughly half of all European, Asian and native American mtDNAs. |
|
|
|
European mtDNAs can be subdivided into two groups according to the presence (25%) or absence (75%) of the DdeI site at bp10394. About 99% of the European mtDNAs can subsumed within 9 mtDNA haplogroups, designated H, I, J, K, T, U, V, W and X (Torroni et al. 1996), and these haplogroups can be further subsumed within four mtDNA haplogroup clusters, HV, UK, TJ and WIX (Richards et al. 1998). In Asia, 77% of all mtDNA are encompassed within a super-haplogroup M defined by a DdeI site gain at bp 10394 and an AluI site gain at bp 10397 (Ballinger et al. 1992a, Torroni et al. 1993, Chen et al. 1995b, Wallace 1995). Haplogroup M is subdivided into smaller sub-haplogroups designated C, D, G and E. Most of the remaining Asian mtDNA are encompassed by haplogroups A, B and F (Torroni et al. 1994). Essentially all Native American mtDNAs fall into haplogroups A, B, C and D (Torroni & Wallace 1994). |
|
|
|
African mtDNA haplogroups |
|
|
|
L1 (L1a, L1b, L1c) - Populations carrying L1 are the descendants of the KhoiSan speaking peoples |
|
|
|
L2 - This haplogroup is east African and it is related with the Bantu expansion |
|
|
|
L2a, the most common clade (62% of the total L2), is the only one widespread all over Africa. Not surprisingly, it is also the one more associated with the Bantu and their expansion |
|
|
|
L2b and L2c are more concentrated in western Africa (particularly Senegal) and are virtually absent in eastern Africans, in Biakaand Mbuti Pygmies, and is rare in southern Africans. It is common in some Senegalese populations. |
|
|
|
L2d is the rarest, but it also appears mainly restricted to western Africa (particularly Western Sahara and Mauritania/Senegal). |
|
|
|
L3 (L3b, L3d, L3e)- This haplogroup is east African and it is related with the Bantu expansion |
|
|
|
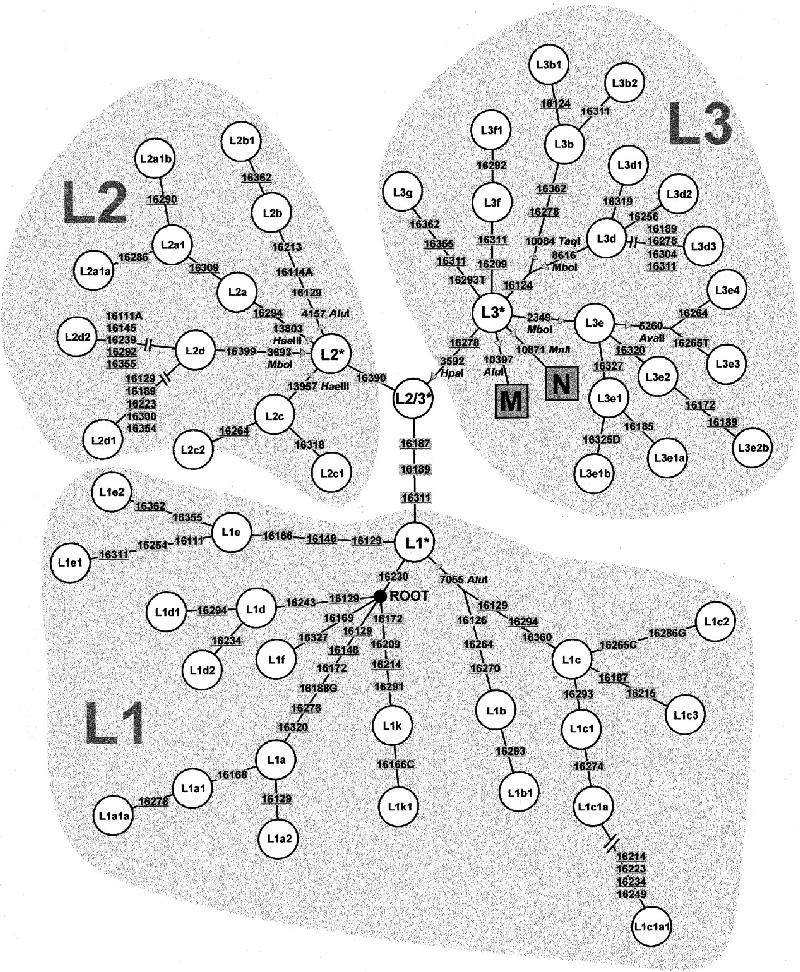
L3* (also known as N/M/L3) - It is particularly important to make it quite clear (particularly to Neo-Nazis like "Refuting Racial Myths") that this group is specific to sub-Saharan Africa, since it is present in a great deal of the European populations including the Finns [Passarino et al. (2002) even found L2 lineages in a Norwegian sample, so the presence of African markers introduced in Nordic populations during the Neolithic shouldn't be a surprise]. This African haplogroup, mostly introduced in Europe during the Neolithic (as explained in Gonzalez et al. 2003) is distinguished from the Eurasian haplogroups M and N at the nucleotide positions 10400 and 10873, respectively (Quintana-Murci et al. 1999). |
|
|
|
 |
|
|
|
This figure (click on the picture for a detailed view) shows quite clearly how deeply N/M/L3 is related to other sub-saharan haplogroups. [Source] |
|
|
|
|
|
Asian and Amerindian mtDNA haplogroups |
|
|
|
M (comprises C, D, E, G) - these 4 haplogroups represent about 77% of the Asian mtDNA |
|
|
|
A, B, F - these haplogroups represent about 23% of the Asian mtDNA |
|
|
|
A, B, C, D - most Amerindian mtDNA fall within these four haplogroups |
|
|
|
M1 - This haplogroup is probably not Negroid. Originally it was given an African origin due to the presence of the subclade M1 in Eastern Africa. However, a posterior return from Asia to Africa of these lineages is a more plausible explanation because the genetic diversity of M is much greater in India than in Ethiopia. In fact, M1 could be a branch of the Indian cluster M as ancestral motifs of the African M1 are found in M*, M3 and M4 Indian subclusters. This supposed Indian expansion to the west also reached northern areas, since evolved representatives of M4 have also been detected in Central Asia. We may consider the upper bound for this return to Africa 25.000-47.000 yr BP, the age calculated for M1 in Eastern Africa. |
|
|
|
European mtDNA haplogroups |
|
|
|
About 99% of the European mtDNA haplogroups fall within haplogroups H, I, J, K, T, U, V, W and X (Torroni et al. 1996). Haplogroup U6, present in the Portuguese population, is of North African (Caucasoid) origin. |
|
|
|
Sources:
- Dr. Mark Joblin - Y Chromosome analysis
- James R Hull - Hull Surname DNA Study
- Dr Emmeline Hill - Inside Ireland
- Dr. Peter Foster - Mitochondrial DNA survey
- Mitochondrial DNA sequence variation in patients with sensorineural hearing impairment and in the Finnish population
- Rousser et al. (2000) - Y Chromosomal Diversity in Europe
- Brehm et al. (2001) - Mitochondrial Portrait of the Cabo Verde Archipelago
- Richards et al. (2000) - Tracing European founder lineages in the Near Eastern mtDNA pool
- Passarino et al. (2002) - Different genetic components in the Norwegian population revealed by the analysis of mtDNA and Y chromosome polymorphisms
- Major genomic mitochondrial lineages delineate early Human expansions
- Torroni et al. (2001) - Do the Four Clades of the mtDNA Haplogroup L2 Evolve at Different Rates?
- Salas et al. (2002) - The Making of the African mtDNA landscape |
|
|
|
Refuting Kemp |
|






